Evaluation of tools for identifying large copy number variations from ultra-low-coverage whole-genome sequencing data | BMC Genomics | Full Text

Bioinformatics Applications Note Genome Analysis Control-free Calling of Copy Number Alterations in Deep-sequencing Data Using Gc-content Normalization | Semantic Scholar
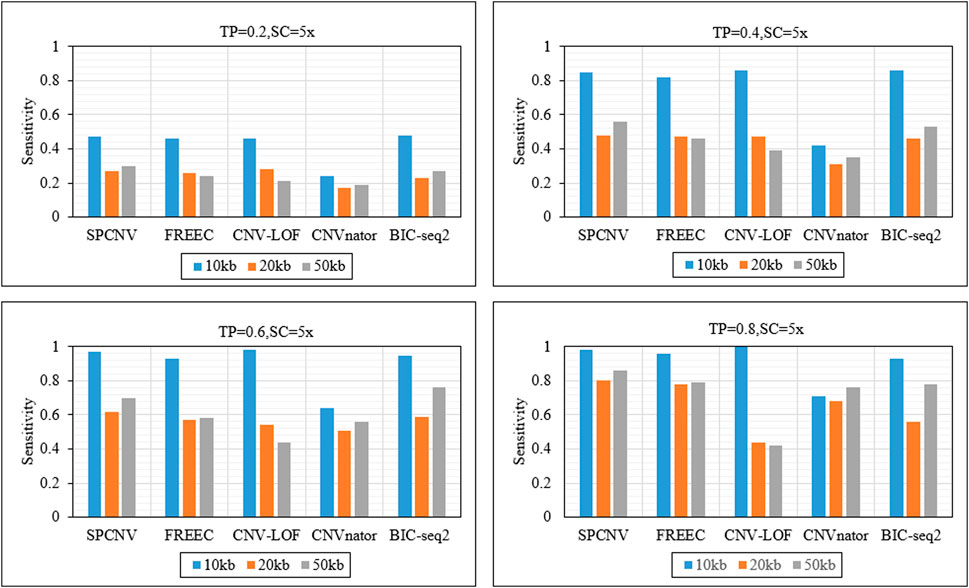
Frontiers | A shortest path-based approach for copy number variation detection from next-generation sequencing data
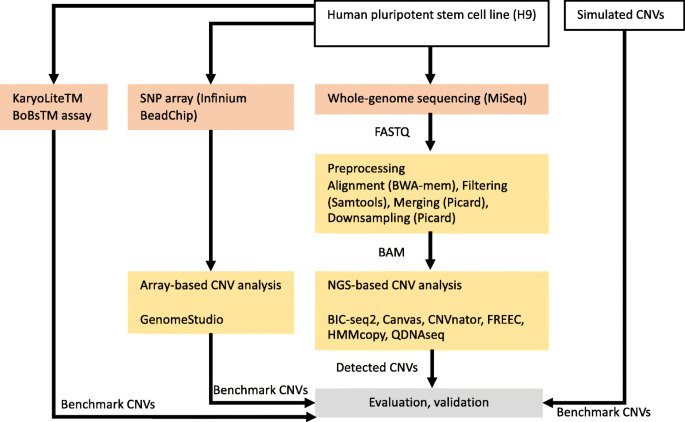
Evaluation of tools for identifying large copy number variations from ultra-low-coverage whole-genome sequencing data
![PDF] Copy number analysis of whole-genome data using BIC-seq2 and its application to detection of cancer susceptibility variants | Semantic Scholar PDF] Copy number analysis of whole-genome data using BIC-seq2 and its application to detection of cancer susceptibility variants | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/aa261b192f8eef42d0724a397ee0515e738b588d/6-Figure3-1.png)
PDF] Copy number analysis of whole-genome data using BIC-seq2 and its application to detection of cancer susceptibility variants | Semantic Scholar
![PDF] Copy number analysis of whole-genome data using BIC-seq2 and its application to detection of cancer susceptibility variants | Semantic Scholar PDF] Copy number analysis of whole-genome data using BIC-seq2 and its application to detection of cancer susceptibility variants | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/aa261b192f8eef42d0724a397ee0515e738b588d/7-Figure4-1.png)
PDF] Copy number analysis of whole-genome data using BIC-seq2 and its application to detection of cancer susceptibility variants | Semantic Scholar

PDF) Copy number analysis of whole-genome data using BIC-seq2 and its application to detection of cancer susceptibility variants
![PDF] dpGMM: A Dirichlet Process Gaussian Mixture Model for Copy Number Variation Detection in Low-Coverage Whole-Genome Sequencing Data | Semantic Scholar PDF] dpGMM: A Dirichlet Process Gaussian Mixture Model for Copy Number Variation Detection in Low-Coverage Whole-Genome Sequencing Data | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/67599f97fcb1685769db1734a10ad2519b8ef1c2/10-Table3-1.png)
PDF] dpGMM: A Dirichlet Process Gaussian Mixture Model for Copy Number Variation Detection in Low-Coverage Whole-Genome Sequencing Data | Semantic Scholar
![PDF] Copy number analysis of whole-genome data using BIC-seq2 and its application to detection of cancer susceptibility variants | Semantic Scholar PDF] Copy number analysis of whole-genome data using BIC-seq2 and its application to detection of cancer susceptibility variants | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/aa261b192f8eef42d0724a397ee0515e738b588d/10-Table1-1.png)
PDF] Copy number analysis of whole-genome data using BIC-seq2 and its application to detection of cancer susceptibility variants | Semantic Scholar
![PDF] Copy number analysis of whole-genome data using BIC-seq2 and its application to detection of cancer susceptibility variants | Semantic Scholar PDF] Copy number analysis of whole-genome data using BIC-seq2 and its application to detection of cancer susceptibility variants | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/aa261b192f8eef42d0724a397ee0515e738b588d/4-Figure1-1.png)
PDF] Copy number analysis of whole-genome data using BIC-seq2 and its application to detection of cancer susceptibility variants | Semantic Scholar
![PDF] WisecondorX: improved copy number detection for routine shallow whole-genome sequencing | Semantic Scholar PDF] WisecondorX: improved copy number detection for routine shallow whole-genome sequencing | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/7ba883a32a465c6790a2ac06aed46b658db5d38f/6-Figure4-1.png)
PDF] WisecondorX: improved copy number detection for routine shallow whole-genome sequencing | Semantic Scholar
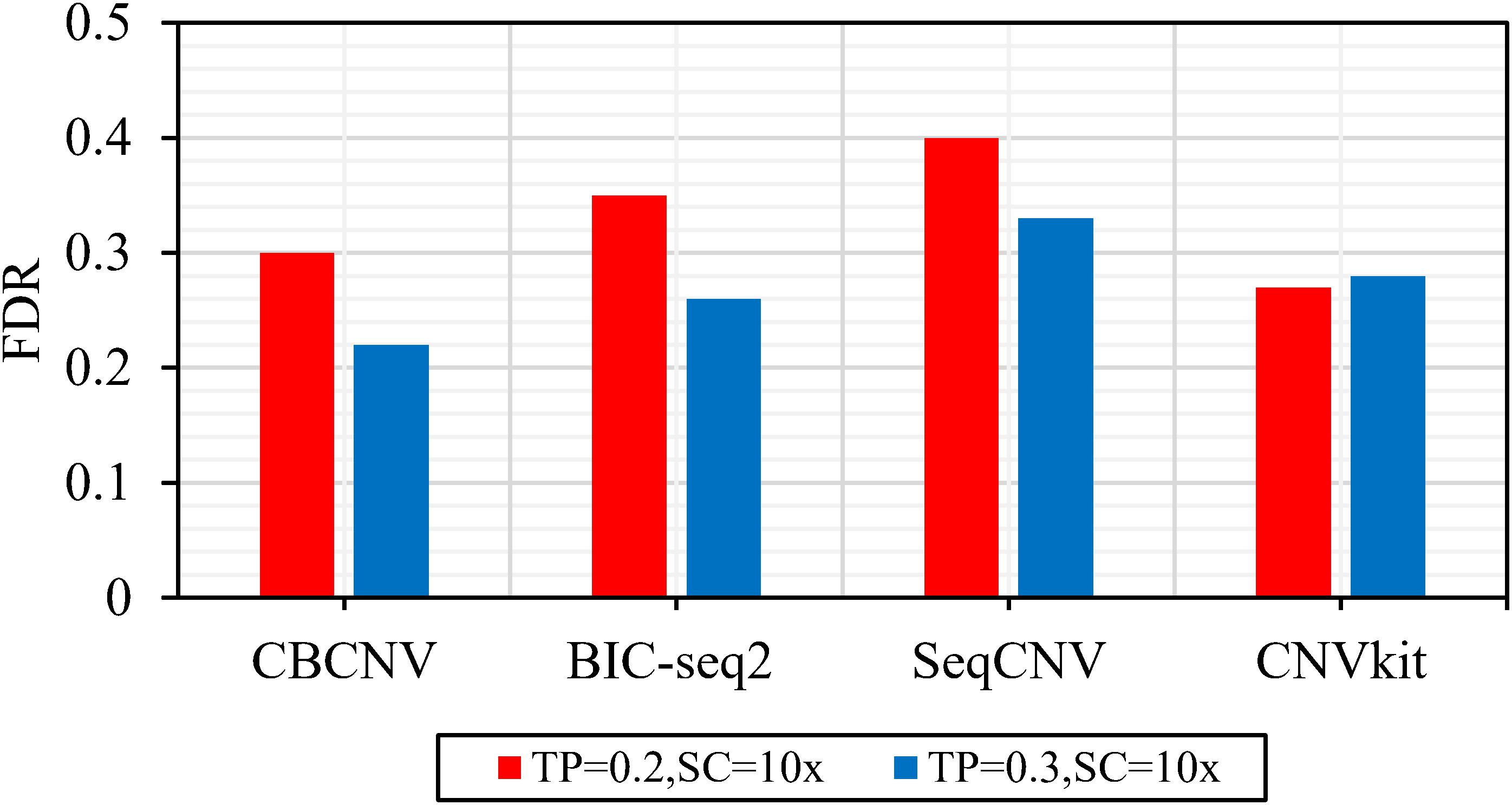
Frontiers | A Cluster-Based Approach for the Discovery of Copy Number Variations From Next-Generation Sequencing Data