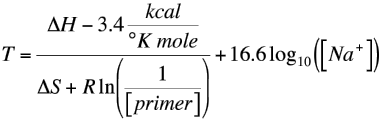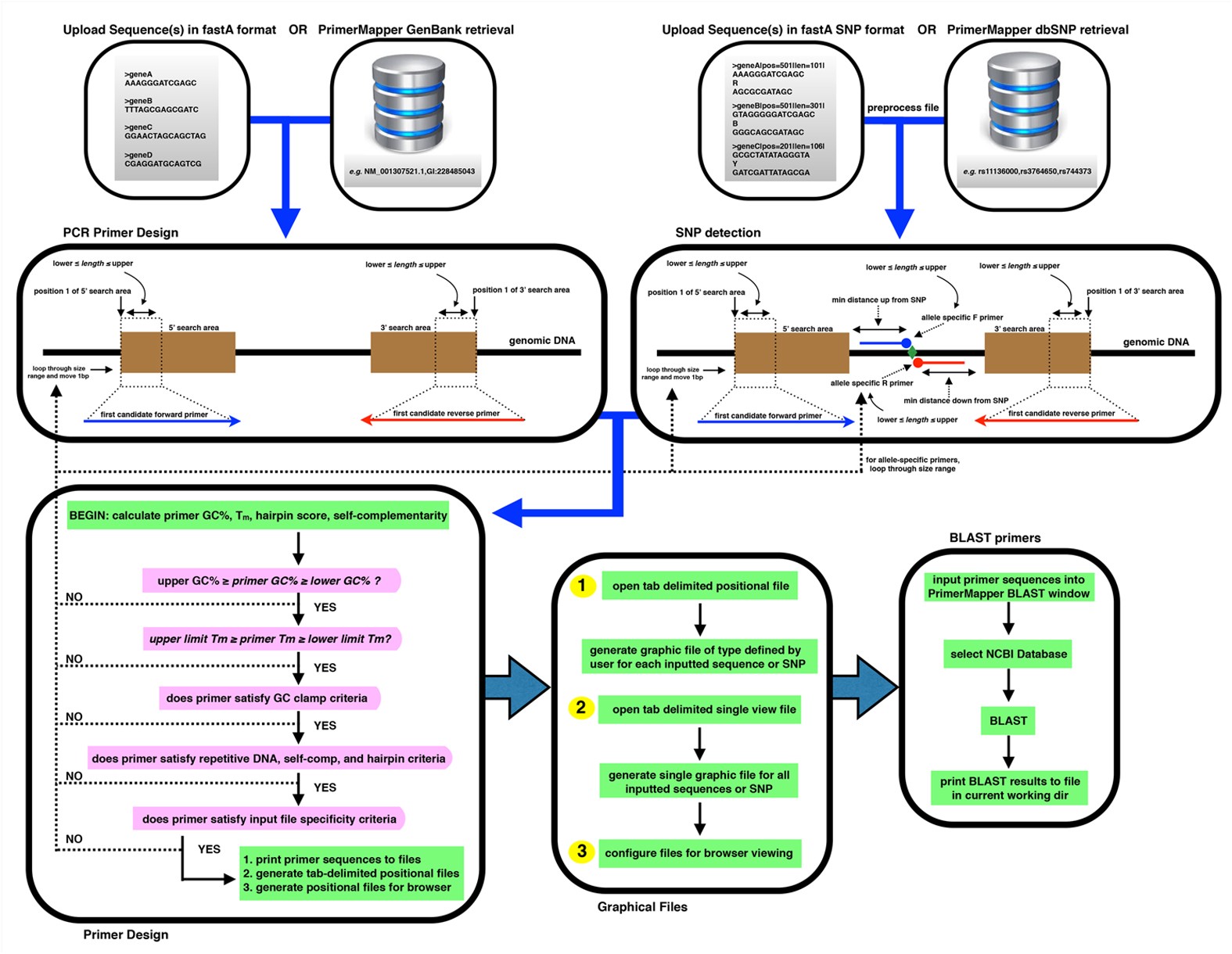
PrimerMapper: high throughput primer design and graphical assembly for PCR and SNP detection | Scientific Reports
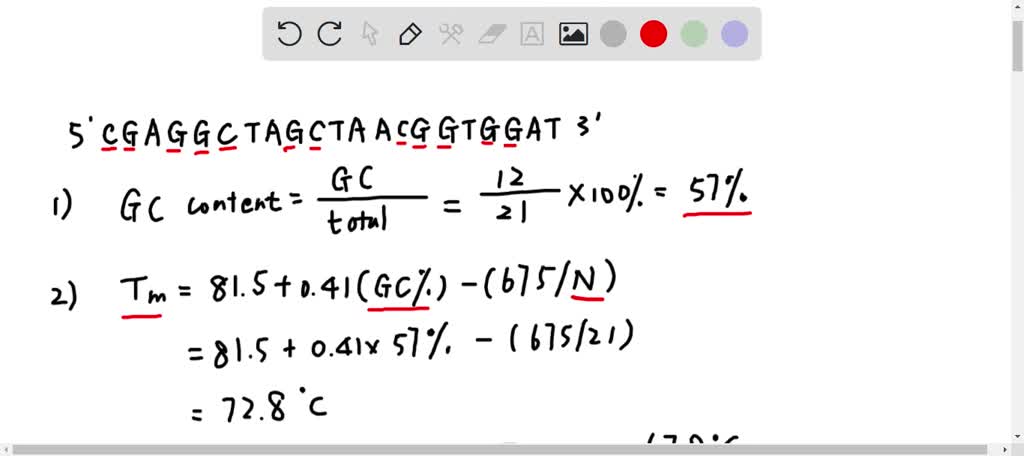
SOLVED: A widely used method for calculating the annealing temperature for a primer used in PCR is 5 degrees below the Tm (ºC), which is computed by this equation 81.5 + 0.41 (%

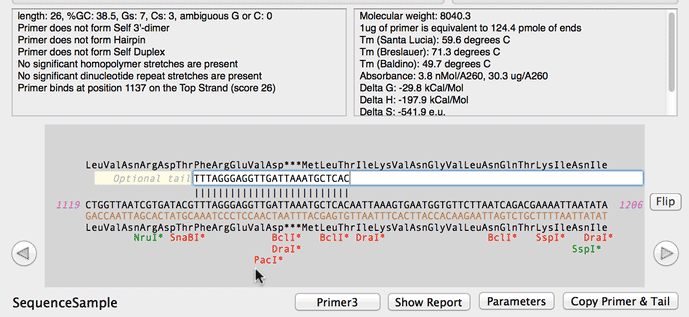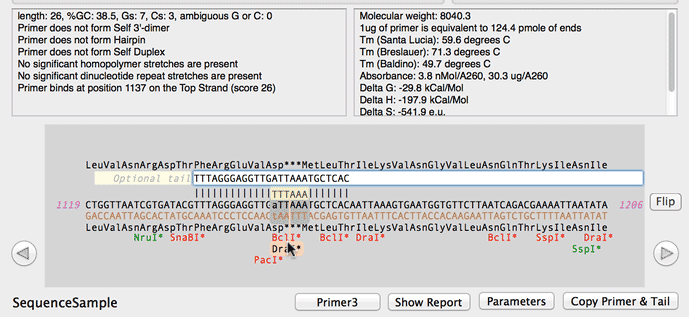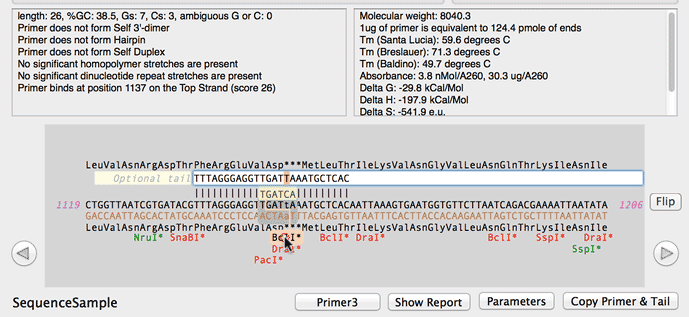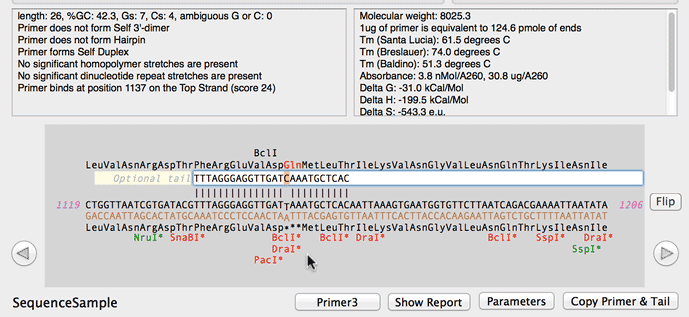
Online oligo-nucleotide calculator entry and main calculator screen... | Download Scientific Diagram